# CCPi-Regularisation Toolkit ([Software X paper](https://www.sciencedirect.com/science/article/pii/S2352711018301912))
| Master | Development | Anaconda binaries |
|--------|-------------|-------------------|
| [](https://anvil.softeng-support.ac.uk/jenkins/job/CILsingle/job/CCPi-Regularisation-Toolkit/) | [](https://anvil.softeng-support.ac.uk/jenkins/job/CILsingle/job/CCPi-Regularisation-Toolkit-dev/) |   [ ](https://anaconda.org/ccpi/ccpi-regulariser) |
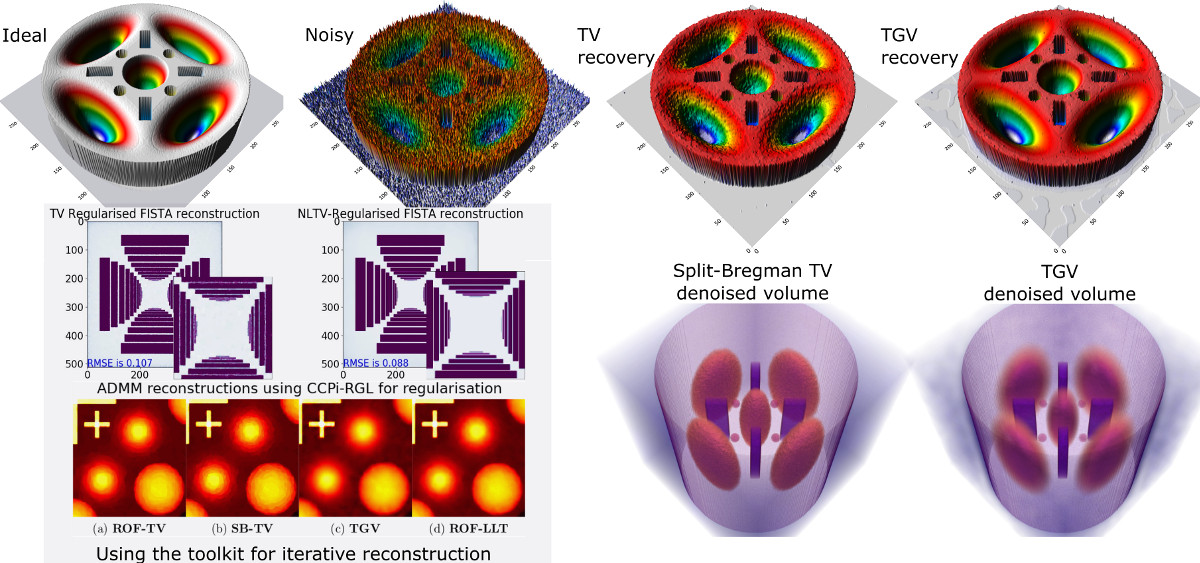
**Iterative image reconstruction (IIR) methods frequently require regularisation to ensure convergence and make inverse problem well-posed. The CCPi-Regularisation Toolkit (CCPi-RGL) toolkit provides a set of 2D/3D regularisation strategies to guarantee a better performance of IIR methods (higher SNR and resolution). The regularisation modules for scalar and vectorial datasets are based on the [proximal operator](https://en.wikipedia.org/wiki/Proximal_operator) framework and can be used with [proximal splitting algorithms](https://en.wikipedia.org/wiki/Proximal_gradient_method), such as PDHG, Douglas-Rachford, ADMM, FISTA and [others](https://arxiv.org/abs/0912.3522). While the main target for CCPi-RGL is [tomographic image reconstruction](https://github.com/dkazanc/ToMoBAR), the toolkit can be used for image denoising and inpaiting problems. The core modules are written in C-OMP and CUDA languages and wrappers for Matlab and Python are provided.**
## Prerequisites:
* [MATLAB](www.mathworks.com/products/matlab/) OR
* Python (tested ver. 3.5/2.7); Cython
* C compilers
* nvcc (CUDA SDK) compilers
## Package modules:
### Single-channel (scalar):
1. Rudin-Osher-Fatemi (ROF) Total Variation (explicit PDE minimisation scheme) **2D/3D CPU/GPU** (Ref. *1*)
2. Fast-Gradient-Projection (FGP) Total Variation **2D/3D CPU/GPU** (Ref. *2*)
3. Split-Bregman (SB) Total Variation **2D/3D CPU/GPU** (Ref. *5*)
4. Primal-Dual (PD) Total Variation **2D/3D CPU/GPU** (Ref. *13*)
5. Total Generalised Variation (TGV) model for higher-order regularisation **2D/3D CPU/GPU** (Ref. *6,13*)
6. Linear and nonlinear diffusion (explicit PDE minimisation scheme) **2D/3D CPU/GPU** (Ref. *8*)
7. Anisotropic Fourth-Order Diffusion (explicit PDE minimisation) **2D/3D CPU/GPU** (Ref. *9*)
8. A joint ROF-LLT (Lysaker-Lundervold-Tai) model for higher-order regularisation **2D/3D CPU/GPU** (Ref. *10,11*)
9. Nonlocal Total Variation regularisation (GS fixed point iteration) **2D CPU/GPU** (Ref. *12*)
### Multi-channel (vectorial):
1. Fast-Gradient-Projection (FGP) Directional Total Variation **2D/3D CPU/GPU** (Ref. *3,4,2*)
2. Total Nuclear Variation (TNV) penalty **2D+channels CPU** (Ref. *7*)
### Inpainting:
1. Linear and nonlinear diffusion (explicit PDE minimisation scheme) **2D/3D CPU** (Ref. *8*)
2. Iterative nonlocal vertical marching method **2D CPU**
## Installation:
The package comes as a [CMake](https://cmake.org) project
and additional wrappers for Python and Matlab.
To install precompiled binaries, you need `conda` and install from `ccpi` channel using :
```
conda install ccpi-regulariser -c ccpi -c conda-forge
```
In order to compile C/C++ sources and additional wrappers from source code for numpy 1.12 and python 3.6, the recommended way is:
```
git clone https://github.com/vais-ral/CCPi-Regularisation-Toolkit
cd CCPi-Regularisation-Toolkit
export CCPI_BUILD_ARGS="--numpy 1.12 --python 3.6"
build/jenkins-build.sh
```
this will install `conda build` environment and compiles C/C++ and Python wrappers and performs basic tests for environment with python 3.6 and numpy 1.12.
### CMake
If you want to build directly using cmake, install CMake (v.>=3) to configure it. Additionally you will need a C compiler, `make` (on linux) and CUDA SDK where available. The toolkit may be used directly from C/C++ as it is compiled as a shared library (check-out the include files in `Core` for this)
1. Clone this repository to a directory, i.e. `CCPi-Regularisation-Toolkit`,
2. create a build directory.
3. Issue `cmake` to configure (or `cmake-gui`, or `ccmake`, or `cmake3`). Use additional flags to fine tune the configuration.
Flags used during configuration
| CMake flag | type | meaning |
|:---|:----|:----|
| `BUILD_PYTHON_WRAPPER` | bool | `ON\|OFF` whether to build the Python wrapper |
| `BUILD_MATLAB_WRAPPER` | bool | `ON\|OFF` whether to build the Matlab wrapper |
| `CMAKE_INSTALL_PREFIX` | path | your favourite install directory |
| `PYTHON_DEST_DIR` | path | python modules install directory (default `${CMAKE_INSTALL_PREFIX}/python`) |
| `MATLAB_DEST_DIR` | path | Matlab modules install directory (default `${CMAKE_INSTALL_PREFIX}/matlab`)|
| `BUILD_CUDA` | bool | `ON\|OFF` whether to build the CUDA regularisers |
| `CONDA_BUILD`| bool | `ON\|OFF` whether it is installed with `setup.py install`|
| `Matlab_ROOT_DIR` | path | Matlab directory|
|`PYTHON_EXECUTABLE` | path | /path/to/python/executable|
Here an example of build on Linux (see also `run.sh` for additional info):
```bash
git clone https://github.com/vais-ral/CCPi-Regularisation-Toolkit.git
cd build
cmake .. -DCONDA_BUILD=OFF -DBUILD_MATLAB_WRAPPER=ON -DBUILD_PYTHON_WRAPPER=ON -DBUILD_CUDA=ON -DCMAKE_BUILD_TYPE=Release -DCMAKE_INSTALL_PREFIX=./install
make install
cd install/python
export LD_LIBRARY_PATH=${LD_LIBRARY_PATH}:../lib
```
### Python
#### Python binaries
Python binaries are distributed via the [ccpi](https://anaconda.org/ccpi/ccpi-regulariser) conda channel. Currently we produce packages for Linux64, Python 2.7, 3.5 and 3.6, NumPy 1.12 and 1.13.
```
conda install ccpi-regulariser -c ccpi -c conda-forge
```
#### Python (conda-build)
```
export CIL_VERSION=19.10.1 (Unix) / set CIL_VERSION=19.10 (Windows)
conda build recipe/ --numpy 1.15 --python 3.7
conda install ccpi-regulariser=${CIL_VERSION} --use-local --force-reinstall # doesn't work?
conda install -c file://${CONDA_PREFIX}/conda-bld/ ccpi-regulariser=${CIL_VERSION} --force-reinstall # try this one
cd demos/
python demo_cpu_regularisers.py # to run CPU demo
python demo_gpu_regularisers.py # to run GPU demo
```
#### Python build
If passed `CONDA_BUILD=ON` the software will be installed by issuing `python setup.py install` which will install in the system python (or whichever other python it's been picked up by CMake at configuration time.)
If passed `CONDA_BUILD=OFF` the software will be installed in the directory pointed by `${PYTHON_DEST_DIR}` which defaults to `${CMAKE_INSTALL_PREFIX}/python`. Therefore this directory should be added to the `PYTHONPATH`.
If Python is not picked by CMake you can provide the additional flag to CMake `-DPYTHON_EXECUTABLE=/path/to/python/executable`.
### Matlab
Matlab wrapper will install in the `${MATLAB_DEST_DIR}` directory, which defaults to `${CMAKE_INSTALL_PREFIX}/matlab`
If Matlab is not picked by CMake, you could add `-DMatlab_ROOT_DIR=`.
#### Linux
Because you've installed the modules in `` you need to instruct Matlab to look in those directories:
```bash
PATH="/path/to/mex/:$PATH" LD_LIBRARY_PATH="/path/to/library:$LD_LIBRARY_PATH" matlab
```
By default `/path/to/mex` is `${CMAKE_INSTALL_PREFIX}/bin` and `/path/to/library/` is `${CMAKE_INSTALL_PREFIX}/lib`
#### Windows
On Windows the `dll` and the mex modules must reside in the same directory. It is sufficient to add the directory at the beginning of the m-file.
```matlab
addpath(/path/to/library);
```
#### Legacy Matlab installation (partly supported, please use Cmake)
```
cd src/Matlab/mex_compile
compileCPU_mex.m % to compile CPU modules
compileGPU_mex.m % to compile GPU modules (see instructions in the file)
```
### References to implemented methods:
1. [Rudin, L.I., Osher, S. and Fatemi, E., 1992. Nonlinear total variation based noise removal algorithms. Physica D: nonlinear phenomena, 60(1-4)](https://www.sciencedirect.com/science/article/pii/016727899290242F)
2. [Beck, A. and Teboulle, M., 2009. Fast gradient-based algorithms for constrained total variation image denoising and deblurring problems. IEEE Transactions on Image Processing, 18(11)](https://doi.org/10.1109/TIP.2009.2028250)
3. [Ehrhardt, M.J. and Betcke, M.M., 2016. Multicontrast MRI reconstruction with structure-guided total variation. SIAM Journal on Imaging Sciences, 9(3)](https://doi.org/10.1137/15M1047325)
4. [Kazantsev, D., Jørgensen, J.S., Andersen, M., Lionheart, W.R., Lee, P.D. and Withers, P.J., 2018. Joint image reconstruction method with correlative multi-channel prior for X-ray spectral computed tomography. Inverse Problems, 34(6)](https://doi.org/10.1088/1361-6420/aaba86) **Results can be reproduced using the following** [SOFTWARE](https://github.com/dkazanc/multi-channel-X-ray-CT)
5. [Goldstein, T. and Osher, S., 2009. The split Bregman method for L1-regularized problems. SIAM journal on imaging sciences, 2(2)](https://doi.org/10.1137/080725891)
6. [Bredies, K., Kunisch, K. and Pock, T., 2010. Total generalized variation. SIAM Journal on Imaging Sciences, 3(3)](https://doi.org/10.1137/090769521)
7. [Duran, J., Moeller, M., Sbert, C. and Cremers, D., 2016. Collaborative total variation: a general framework for vectorial TV models. SIAM Journal on Imaging Sciences, 9(1)](https://doi.org/10.1137/15M102873X)
8. [Black, M.J., Sapiro, G., Marimont, D.H. and Heeger, D., 1998. Robust anisotropic diffusion. IEEE Transactions on image processing, 7(3)](https://doi.org/10.1109/83.661192)
9. [Hajiaboli, M.R., 2011. An anisotropic fourth-order diffusion filter for image noise removal. International Journal of Computer Vision, 92(2)](https://doi.org/10.1007/s11263-010-0330-1)
10. [Lysaker, M., Lundervold, A. and Tai, X.C., 2003. Noise removal using fourth-order partial differential equation with applications to medical magnetic resonance images in space and time. IEEE Transactions on image processing, 12(12)](https://doi.org/10.1109/TIP.2003.819229)
11. [Kazantsev, D., Guo, E., Phillion, A.B., Withers, P.J. and Lee, P.D., 2017. Model-based iterative reconstruction using higher-order regularization of dynamic synchrotron data. Measurement Science and Technology, 28(9)](https://doi.org/10.1088/1361-6501/aa7fa8)
12. [Abderrahim E., Lezoray O. and Bougleux S. 2008. Nonlocal discrete regularization on weighted graphs: a framework for image and manifold processing. IEEE Trans. Image Processing 17(7), pp. 1047-1060.](https://ieeexplore.ieee.org/document/4526700)
13. [Chambolle, A. and Pock, T., 2010. A first-order primal-dual algorithm for convex problems with applications to imaging. Journal of mathematical imaging and vision 40(1)](https://doi.org/10.1007/s10851-010-0251-1)
### References to software (please cite if used):
* [Kazantsev, D., Pasca, E., Turner, M.J. and Withers, P.J., 2019. CCPi-Regularisation toolkit for computed tomographic image reconstruction with proximal splitting algorithms. SoftwareX, 9, pp.317-323.](https://www.sciencedirect.com/science/article/pii/S2352711018301912)
### Applications:
* [TOmographic MOdel-BAsed Reconstruction (ToMoBAR)](https://github.com/dkazanc/ToMoBAR)
* [Joint image reconstruction method with correlative multi-channel prior for X-ray spectral computed tomography (MATLAB code)](https://github.com/dkazanc/multi-channel-X-ray-CT)
### License:
[Apache License, Version 2.0](http://www.apache.org/licenses/LICENSE-2.0)
### Acknowledgments:
CCPi-RGL software is a product of the [CCPi](https://www.ccpi.ac.uk/) group and STFC SCD software developers. Any relevant questions/comments can be e-mailed to Daniil Kazantsev at dkazanc@hotmail.com

